- Faculty of Biological Sciences
- Research and innovation
Research and innovation
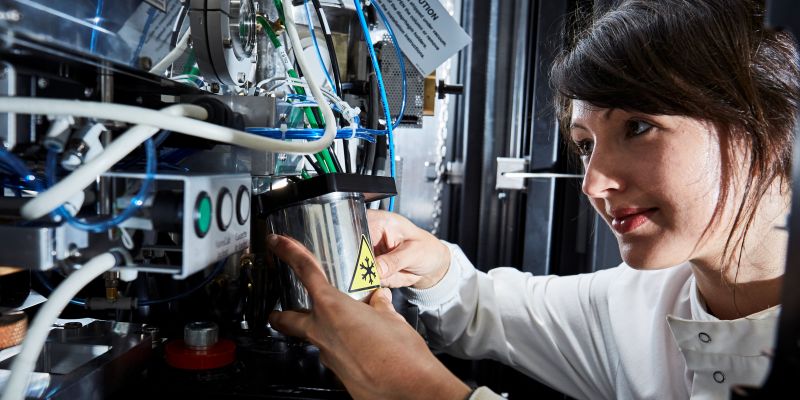
The Faculty of Biological Sciences is a vibrant, interdisciplinary, and collaborative research community, spanning all areas of biological sciences from molecules to landscapes. Our research and innovation pages highlight the strength and diversity of our research teams. Do contact us if you have an interest in working together.
At least 88%
'world leading' or 'internationally excellent'
Submitted research in our two main UoAs – REF 2021
£90m
research laboratories redesign
to enhance collaboration
£17m
Astbury BioStructure Laboratory
For world-leading biological research
Our Schools
We take a collaborative approach to delivering impactful research at the Faculty of Biological Sciences. Each of our schools also has its own research expertise and focus:
Our research
Our faculty has over 130 academic staff, 14 independently funded research fellows and 200+ PhD students working at all levels of biological sciences, from molecules to landscapes. The links below allow you to explore our current research areas across our three schools.
Biotechnology
Biotechnology is a broad discipline in which biological processes, organisms, cells or cellular components are exploited to develop new technologies.
More on BiotechnologyCancer
Cancer research studies the molecular and cellular basis of cancer and the development of target-specific therapeutics.
More on CancerCardiovascular
Our Cardiovascular research is delivered under 6 themes and focuses on the function of the heart in health and disease, with additional activity in the area of skeletal muscle.
More on CardiovascularCell and Organismal Biology
Our Cell and Organismal Biology research aims at determining the cellular and molecular mechanisms underlying both normal cell functions and how they become dysfunctional in diseased cells and organisms.
More on Cell and Organismal BiologyEcology and Evolution
Our Ecological and Evolutionary research ranges from fundamental theory through to directly applied work, at scales ranging from molecular to global and across all the Earth’s biomes.
More on Ecology and EvolutionHeredity, Development and Disease
Our Heredity, Development and Disease research studies a variety of organisms to investigate fundamental processes in animal and human development and disease.
More on Heredity, Development and DiseaseIntegrative Membrane Biology
Within Integrative Membrane Biology research, our multidisciplinary and highly collaborative environment facilitates the study of membrane protein systems from the atomic to the cellular and organismal levels.
More on Integrative Membrane BiologyMicrobiology
Our Microbiology research delivers ground-breaking insights into the fundamental biology of bacteria, viruses and fungi, and has a particular emphasis on understanding in molecular detail the mechanisms by which these agents cause disease and evade treatment.
More on MicrobiologyNeuroscience
Our Neuroscience research seeks to understand how the nervous system, the most complex and highly organised part of the body, is able to generate perception, thoughts and behaviour.
More on NeurosciencePlant Sciences
Plant Science research address fundamental biological mechanisms as related to plants and also addresses the global challenge of maintaining food supplies in the context of sustainable agriculture.
More on Plant SciencesSport and Exercise Sciences
Sport and Exercise Science research has a world-class focus for research excellence in exercise science, building on its distinctive identity in understanding the basic mechanisms, and translational applications of physical activity in health and disease.
More on Sport and Exercise SciencesStructural Biology
Our Structural Biology research aims to understand the structures of biomolecules such as proteins, nucleic acids, carbohydrates and lipids, and the complexes they form.
More on Structural BiologyTissue Engineering
Our Tissue Engineering research focuses on the development of scaffolds and medical devices to regenerate damaged and diseased parts of the body.
More on Tissue EngineeringA new home for teaching and research
The Faculty of Biological Sciences is undergoing a major refurbishment to develop its teaching and research capabilities. Creating highly-flexible state of the art laboratories, offices and study space, the investment will provide an exciting new environment for an outstanding student experience and for large multidisciplinary research teams from across multiple faculties to work together.
Research Centres
Below are some of the cross-faculty research centres within the University that draw together expertise, maximising potential for application of novel approaches to solve scientific problems.
Leeds Omics
Our mission is to engage and unify the critical mass of ‘omics’ researchers at the University of Leeds into one central virtual institute. We have a significant number of active researchers that fit under this “Omics” umbrella.
More on Leeds OmicsThe Astbury Centre
Bringing together researchers from across the University - largely from physics, the biological sciences and chemistry - to allow interdisciplinary approaches to be harnessed to understand the molecular basis of life.
More on The Astbury CentreCentre for Plant Sciences
A centre for excellence in plant cell and molecular biology. The Centre has involvement in many European consortia and an extensive range of international collaborations including with the USA, India, Japan and China.
More on Centre for Plant SciencesInstitute of Medical and Biological Engineering
Focusing on longer lasting joint replacements, tissue sparing interventions and biological scaffolds for tissue regeneration supported by computational and experimental simulation systems for design and pre-clinical testing.
More on Institute of Medical and Biological EngineeringResearch Facilities
Our 10 facilities cover the broad research themes of biophysical characterisation, structural elucidation and cellular visualisation. They provide a full pipeline for preparation and complete characterisation of systems from single molecules, to macro-molecular complexes, to cells.
Research Impact
Our research delivers significant benefits to society and the economy. Read through case studies from some of our leading academics that showcase how we have already delivered a high level of impact across the globe as well as exploring our early future impact news stories.
Biomedicine and Health

Find out how we are tackling challenges within Biomedicine and Health by reading our impact case studies and some of our other research news.
MoreEcology and Conservation

Find out how we are tackling challenges within Ecology and Conservation by reading our impact case studies and some of our other research news.
MoreSport and Exercise Sciences

Find out how we are tackling challenges within Sport and Exercise Science by reading our impact case studies and some of our other research news.
MoreSustainable Agriculture

Find out how we are tackling challenges within Sustainable Agriculture by reading our impact case studies and some of our other research news.
MoreScientists inspire the next generation
Wednesday 10 April 2024
More on Scientists inspire the next generationPostdoctoral Symposium 2024
Thursday 20 June 2024, 09:00 - 17:00 |
More on Postdoctoral Symposium 2024